Welcome to pyIMD’s documentation!¶




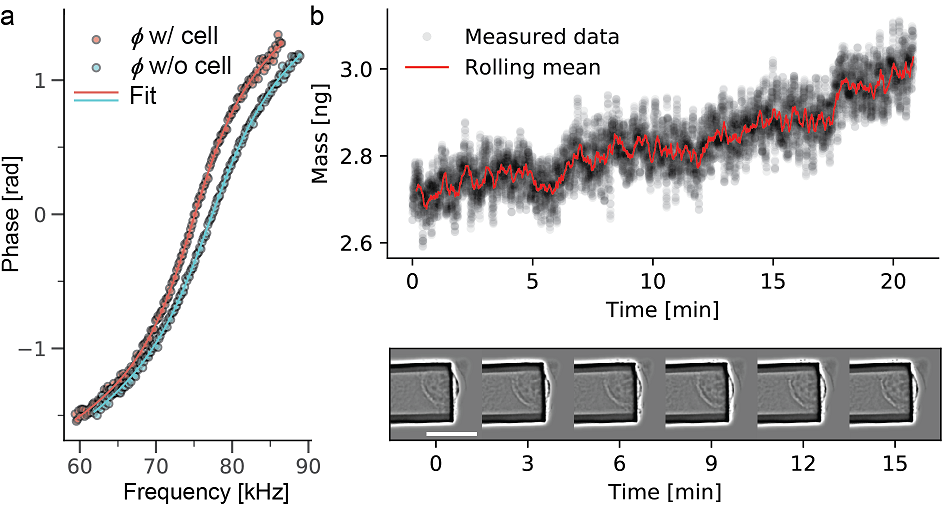
Evolution of mass over time and the corresponding microscopy images are shown for a time span of 20min. The mass data was acquired every 10 ms (data shown in black), overlaid in red is the rolling mean with a window of 1000. Images taken every 3 min over the observed times span. The mammalian cell increases mass steadily..
The total mass of single cells can be accurately monitored in real time under physiological conditions with our recently developed picobalance. It is a powerful tool to investigate crucial processes in biophysics, cell biology or medicine, such as cell mass regulation. However, processing of the raw data can be challenging, as computation is needed to extract the mass and long-term measurements can generate large amounts of data. Here, we introduce the software package pyIMD that automates raw data processing, particularly when investigating non-migrating cells. pyIMD stands for Python inertial mass determination and is implemented using Python >=3.5 and can be used as a command line tool or as a stand-alone version including a graphical user interface.
This documentation of pyIMD describes the API and provides sample data sets as well as sample scripts to run pyIMD from Jupyter or the Python console. It also contains a tutorial about how pyIMD is used with the user interface.